Abstract
Historically, studies of disseminated tumour cells in bone marrow and circulating tumour cells in peripheral blood have provided crucial insights into cancer biology and the metastatic process. More recently, advances in the detection and characterization of circulating tumour DNA (ctDNA) have finally enabled the introduction of liquid biopsy assays into clinical practice. The FDA has already approved several single-gene assays and, more recently, multigene assays to detect genetic alterations in plasma cell-free DNA (cfDNA) for use as companion diagnostics matched to specific molecularly targeted therapies for cancer. These approvals mark a tipping point for the widespread use of liquid biopsy in the clinic, and mostly in patients with advanced-stage cancer. The next frontier for the clinical application of liquid biopsy is likely to be the systemic treatment of patients with ‘ctDNA relapse’, a term we introduce for ctDNA detection prior to imaging-detected relapse after curative-intent therapy for early stage disease. Cancer screening and diagnosis are other potential future applications. In this Perspective, we discuss key issues and gaps in technology, clinical trial methodologies and logistics for the eventual integration of liquid biopsy into the clinical workflow.
Similar content being viewed by others
Introduction
Intratumoural heterogeneity and tumour evolution contribute to treatment failure in patients with cancer1. Over the disease course, these processes are influenced and enhanced by cellular mutability and include described interactions between whole-genome doubling and cancer cell survival2, metabolic and homeostatic mechanisms, microenvironmental factors (for example, hypoxia) and drug selection pressures, including adaptive mutability3,4, all of which influence the abundance and characteristics of tumour cells and products that are shed and can be measured in the bloodstream at different times5. Intratumoural and intertumoural heterogeneity have been elucidated through large collaborative efforts using different omics technologies for primary and metastatic tumour analyses, including analyses by the International Cancer Genome Consortium (ICGC)6, The Cancer Genome Atlas (TCGA)7, the Pan-Cancer Analysis of Whole Genomes Consortium of the ICGC and TCGA8, and the Human Tumor Atlas Network9. This heterogeneity has been further elucidated through single-cell analyses of primary tumours and corresponding metastases10,11, genomics studies of treated and untreated metastatic disease12,13 and investigations of the tumour microenvironment14. Advances in the study of different liquid biopsy analytes over the past two decades have led to the hope that any spatial and temporal heterogeneity in tumour biology could be better tracked by serial blood analyses than by analyses of tissue samples from a primary tumour that might have been excised years earlier and prior to systemic therapy or from a single metastatic lesion, when multiple potentially discordant metastases are present, and thus lead to improvements in patient management and outcomes15,16. In comparison with tumour tissue-based approaches, liquid biopsy is less invasive and samples are more easily obtainable throughout the course of disease, thus complementing the development of dynamic molecular imaging assessments17. These liquid biopsy analytes include circulating tumour cells (CTCs), circulating nucleic acids (including circulating tumour DNA (ctDNA), the tumour-derived fraction of cell-free DNA (cfDNA) in the plasma, as well as cell-free RNAs (mRNAs, long non-coding RNAs and microRNAs), extracellular vesicles, tumour-educated platelets, proteins and metabolites that can be found in a range of bodily fluids5,11. Despite the technological advances, the uptake of liquid biopsy in clinical practice has been slow18.
Herein, we summarize selected advances in the use of CTC and ctDNA technologies, clinical trial methodology and implementation logistics that we believe are necessary for liquid biopsy to fulfil its potential to transform the practice of oncology. We discuss the applications of liquid biopsy for treatment selection and disease monitoring in patients with early stage and advanced-stage disease as well as some contemporary data on the use of liquid biopsy for early cancer diagnosis (Fig. 1), focusing on CTCs and ctDNA (Table 1); however, we do so in the context of a Perspective, providing our views on this field mainly for solid tumours. Many excellent and comprehensive reviews on liquid biopsy and its application in specific cancer types are available in this journal19,20 and elsewhere5,16,21,22.
A single blood sample can contain a range of cell types and cell products emanating from multiple tumour sites around the body. Liquid biopsy assays of these tumour-derived factors can serve several purposes in the management of cancer. (1) Early detection of cancer; liquid biopsy approaches could also be used to further investigate abnormalities detected on imaging examinations such as mammography or lung CT. (2) Surveillance for micrometastatic disease following curative-intent treatment of a primary tumour, in order to evaluate the risk of disease recurrence and enable timely management of recurrent disease, if needed. (3) Guiding the selection of the most appropriate treatment and/or monitoring treatment responses in patients with overt metastatic disease through dynamic characterization of changes in tumour burden and disease biology. CTC, circulating tumour cell; ctDNA, circulating tumour DNA; TEPs, tumour-educated platelets.
Stages of assay development
The clinical adoption of a liquid biopsy assay is achieved through three distinct steps, each of which has several important requirements that must be met (Fig. 2). The first step involves the development and validation of the assay. According to the Evaluation of Genomic Applications in Practice and Prevention (EGAPP) initiative, this step includes three critical components: analytical validation, clinical validation and demonstration of clinical utility23. Analytical validity refers to the ability of an assay to reliably and accurately measure the analyte of interest and is evaluated according to the assay’s sensitivity, specificity, reliability and robustness. Clinical validity refers to the ability of an assay to reliably and accurately measure the clinical feature of interest and is evaluated on the basis of clinical sensitivity, specificity, and positive and negative predictive values. Clinical utility is indicated by evidence of improved clinical outcomes (which might include clinical efficacy or reduced toxicity) compared with standard methods when the novel assay is used to direct patient management. These three criteria need to be evaluated based on available evidence in the literature in order to determine whether an assay is likely to achieve regulatory clearance and/or reimbursement.
This roadmap includes three steps. Step 1 encompasses the development and validation of the assay, including demonstration of its clinical feasibility, reproducibility and value. Step 2 involves regulatory approval, incorporation into guidelines for diagnosing and treating cancer, and assay reimbursement, which is dependent on the evidence obtained during step 1 and often additional analyses of cost-effectiveness. Finally, step 3 relates to incorporation of the assay into the clinical workflow, with associated resources and logistical and educational requirements.
Liquid biopsy assays are often developed based on an analysis of whole-genome sequencing (WGS) or whole-exome sequencing (WES) data from tumour tissue samples, which can then be refined to large and/or customized gene panels. One well-validated example is the FDA-authorized Integrated Mutation Profiling of Actionable Cancer Targets gene panel developed at the Memorial Sloan Kettering Cancer Center (MSK-IMPACT), which was initially developed as a hybrid capture-based next-generation sequencing (NGS) assay for targeted deep sequencing of key cancer genes in formalin-fixed, paraffin-embedded (FFPE) tumour specimens24. This assay was used to study DNA from >10,000 tumour specimens and patient-matched germline DNA derived from peripheral blood in order to identify clinically relevant mutations and mutation signatures, as well as novel non-coding alterations, shared between common and rare tumour types25. This panel has now been expanded to interrogate 468 cancer-related genes, again through the analysis of tumour-derived and matched germline DNA samples performed in a Clinical Laboratory Improvement Amendments (CLIA)-certified laboratory26, and is approved by the FDA as a tumour profiling test to provide information on somatic alterations and microsatellite instability (MSI) for use by qualified health-care professionals in accordance with professional guidelines27.
This type of panel discovery process can be aided by open-access tools such as OncoPrint, which enables genetic alterations such as somatic mutations, copy number alterations (CNAs) and changes in mRNA expression to be visualized as a heatmap across datasets28. This tool is available for use in the cBioPortal for Cancer Genomics, another open-access resource for exploring multidimensional cancer genomics datasets29. OncoKB is another publicly available knowledge base and tool that curates and annotates the biological, prognostic and predictive relevance of somatic molecular alterations associated with cancer30.
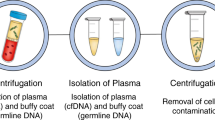
In addition to the tissue-based MSK-IMPACT assay, the MSK-ACCESS (Analysis of Circulating Cell-free DNA to Evaluate Somatic Status) assay has been developed by researchers at the same institution for broad coverage of cancer-associated genes using deep sequencing of plasma cfDNA. This liquid biopsy assay was approved for use in the identification of molecular and cellular tumour markers by the New York State Department of Health in 2019 (ref.31). Importantly, through paired analyses of both the plasma and buffy coat from the same blood samples, cfDNA profiles can be compared with the genomic DNA of white blood cells collected from the same patient to account for germline variants or potential clonal haematopoiesis (which is discussed in a later section of this article) and thus improve the accuracy of ctDNA detection32. We wish to emphasize, however, that the development, validation and subsequent regulatory filing of an in-house multigene assay, such as MSK-IMPACT or MSK-ACCESS, is arduous, labour-intensive and time-consuming, requiring substantial laboratory and clinical expertise, patient sample accrual and financial investment, making it an unrealistic goal for most laboratories to achieve.
A key aspect of assay development relates to changes in assay performance attributable to preanalytical issues, such as the type of specimen analysed, specific procedures of sample collection, handling, processing and storage, and patient-related factors. Indeed, the standardization of preanalytical variables is now considered an important part of the assay development process. The format used for reporting and interpretation of the assay results is another equally important area, especially for genomic data. In this regard, assay developers have a responsibility to produce evidence-based annotations of the results so that clinicians and multidisciplinary tumour boards can provide evidence-based treatment recommendations based on the resultant report. Several efforts have been made to facilitate this process by ranking genomic aberrations according to their relevance to precision medicine, such as the European Society of Medical Oncology Scale of Actionability of Molecular Targets33 or the aforementioned OncoKB knowledge base. Indeed, assay developers have attempted to provide annotations regarding the actionability of mutational events detected by NGS. Actionability — which is often poorly defined, documented or demonstratable with a high level of evidence — is of particular importance in the clinic, where busy physicians are often required to make rapid and irrevocable therapeutic choices based on limited information. However, when different platforms providing such functionality have been compared using the same sequencing data, they have been shown to vary with regard to determination of actionability34. Increasingly large clinical datasets correlating cancer-associated alterations with therapeutic outcomes for specific drugs will accumulate rapidly in the coming years, and novel methods for harmonizing clinically relevant data from disparate databases are likely to improve the clinical utility of these knowledge bases35.
Once a liquid biopsy assay with convincing analytical validity, clinical validity and clinical utility has been developed, the second step towards clinical integration includes regulatory clearance of the assay for a particular indication, incorporation into treatment guidelines and, eventually, assay reimbursement by payers. This second step involves not only the evaluation of data on assay performance, as previously detailed for the first step in assay development but can also include cost-effectiveness analyses (Fig. 2). The regulatory approval and reimbursement pathways are somewhat different in the USA, Europe and Asia, which together with differences in the structure and funding of health-care systems in these regions at least partly explains variations in the availability of and access to liquid biopsy assays36.
The third and final step in the clinical development of an assay involves incorporation into the clinical workflow. This process requires substantial local investment of time and money in developing the human and laboratory resources and procedures to ensure effective application of the test (Fig. 2) and is further detailed in a subsequent section of this article (Clinical integration of liquid biopsy). Importantly, when an institution is considering in-house use of a commercial liquid biopsy assay (for example, the Oncomine cell-free nucleic acid assays, the AVENIO ctDNA targeted kit, or the QIAseq targeted DNA panels), on-site validation should be performed before application of the assay as a companion diagnostic in a clinical trial or in routine clinical practice.
Preanalytical issues with liquid biopsy
For ctDNA assays, the use of plasma rather than serum is generally preferred because the latter contains higher levels of non-tumour cfDNA generated mostly from leukocyte lysis that occurs with blood clotting during serum preparation. The dilution of ctDNA by leukocyte cfDNA can adversely affect detection of the former, especially ctDNA harbouring low allele fraction mutations37.
Concerning the optimal blood collection tubes for ctDNA detection in plasma, multiple studies have compared tubes containing di-potassium or tri-potassium ethylenediaminetetraacetic acid (EDTA) with those containing heparin or citrate to prevent clotting or with commercial tubes that contain preservative reagents for leukocyte stabilization to prevent lysis38,39,40,41. Currently, at least nine types of collection tube specifically designed for the preservation of cell-free nucleic acids (including cell-free methylated DNA) are commercially available, in which blood samples can be shipped and stored at 18–25 °C (room temperature) for plasma processing 3 to 7 days after collection41,42, in accordance with the manufacturers’ instructions. However, cfDNA-specific guidelines developed by the National Cancer Institute (NCI) Biorepositories and Biospecimen Research Branch Biospecimen Evidence-Based Practices (BEBP) recommend shorter durations of room temperature storage for both EDTA (2–4 h) and preservative tubes (up to 3 days) prior to prescribed methods of plasma isolation and storage at −80 °C (ref.43).
Few data are available on the effects of patient-related factors such as pregnancy, smoking, exercise and various non-malignant conditions that might affect cfDNA levels in blood44. Thus, the associations between patient-related factors and performance of specific ctDNA assays should be carefully explored in prospective studies.
For CTC assays, the use of CellSave tubes provides a window of up to 96 hours for sample processing using the FDA-approved CELLSEARCH platform; however, the cells collected in these tubes are fixed and, therefore, not viable45. Thus, these tubes can be used for downstream applications related to DNA, but not RNA or functional analyses. Currently, tubes that enable the reliable long-term storage of viable CTCs for downstream large-scale omics applications or functional assays, including the generation of cell cultures or mouse xenograft models for use in drug testing, remain under development46. Consequently, the benefits of CTCs over ctDNA in terms of extending analysis beyond genomics (Table 1) can only be realized today in the context of single-centre studies involving small numbers of samples47,48. The development of collection tubes for long-term storage of viable CTCs is a crucial unmet need for the evaluation of the full potential of these analytes in multicentre trials. The typically low numbers of CTCs present in blood samples also hampers testing of the clinical utility of CTC-based functional assays or omics analyses. Leukapheresis has been suggested as a potential way to address this issue49; however, it is questionable whether this approach is safe, can be applied to the majority of patients with cancer and can be performed beyond specialized centres. Other methods to increase CTC yield are under development, such as continuous CTC capture via indwelling catheters or microfluidic chips50.
Guidelines have been proposed regarding the preanalytical conditions for cfDNA analyses, including issues of quality control in blood collection, plasma preparation, cfDNA extraction with documentation of the DNA extraction kit and quantification method used (variations in extraction yield exist among different commercial kits, with yield also dependent on fragment size51) and cfDNA storage, blood sample transport conditions, and biological and demographic factors52. In addition to the NCI BEBP guidelines43, initiatives such as CANCER-ID53 and the European Liquid Biopsy Society (ELBS) in Europe and the Blood Profiling Atlas in Cancer (BloodPAC)54 in the USA aim to define preanalytical standards and best practices for liquid biopsy assay development in order to enable the optimal incorporation of such assays into clinical care20,50. Moreover, the Biomarker Consortium of the US Foundation for the National Institutes of Health is working on identifying and validating reference materials for use in quality control across different ctDNA assays55.
Liquid biopsy in advanced-stage disease
In patients with advanced-stage cancers, liquid biopsy assays can be used for treatment selection, monitoring of treatment efficacy and identifying the most appropriate subsequent treatment when resistance develops. In the following sections, we discuss each of these aspects, focusing on prominent examples to date.
Selecting the right treatment
Detection of druggable targets
PCR-based cfDNA assays for oncogenic driver variants of genes such as EGFR in non-small-cell lung cancer (NSCLC) and KRAS in colorectal cancer (CRC) have high specificity (mean 96%) but only moderate sensitivity (mean 66%) compared with tumour tissue assays, which remain the gold standard44,56; however, use of droplet digital PCR (ddPCR) assays increases the level of sensitivity57. For example, in the PRODIGE-14 trial58, a specific KRAS mutation was detectable using ddPCR with over 90% sensitivity in 42 (91%) of 46 patients with the same KRAS mutation detected in tumour tissue. A PCR-based assay (cobas EGFR mutation test v2) was the first liquid biopsy assay approved by the FDA, as a companion diagnostic test to screen for EGFR mutations in plasma cfDNA from patients with advanced-stage NSCLC who are being considered for treatment with the EGFR tyrosine kinase inhibitor (TKI) erlotinib; if no EGFR exon 19 deletion and/or no exon 21 L858R mutation is detected in cfDNA, EGFR status should subsequently be determined using a FFPE tumour tissue sample44,59.
On the basis of data from the phase III SOLAR-1 trial60, a PCR-based assay (the therascreen PIK3CA RGQ PCR kit) has also been approved by the FDA as a companion diagnostic for the detection of PIK3CA mutations in tumour tissue or plasma from patients with advanced-stage hormone receptor (HR)+/HER2− breast cancer to determine their eligibility for treatment with the PI3Kα inhibitor alpelisib in combination with fulvestrant61. As with the EGFR assay, failure to detect a PIK3CA mutation in cfDNA warrants reflex testing of tumour tissue.
In addition to PCR-based single-gene or multigene assays62, high-throughput NGS-based multigene liquid biopsy tests that can detect a range of genomic alterations, including single-nucleotide variants (SNVs), CNAs, fusions and/or insertion or deletions (indels), have been developed. These assays include the following: Archer Reveal ctDNA 28 (28 genes)63; FoundationACT (62–70 genes, incorporating measurement of blood tumour mutational burden (bTMB))64, which was the predecessor to the current FoundationOne Liquid CDx (F1 Liquid CDx) assay that includes >300 genes as well as measurements of bTMB, MSI and tumour fraction values; Guardant360 CDx (54–73 genes, plus detection of MSI, across successive versions of the assay)65,66,67; Inivata InVision (36–37 commonly mutated genes)68; MSK-ACCESS (129 cancer-associated genes, as discussed above); OncoDNA OncoSTRAT&GO (27 genes)69; PGDx elio plasma resolve (33 genes, or 80 genes in the custom PlasmaSelect-R lung panel)70; Resolution Bioscience Resolution ctDx-Lung (21 genes)63; and multiple other assays for simultaneous profiling of gene panels that are commercially available or under development. These multigene liquid biopsy assays have the advantage of enabling broad and informative cancer genotyping, as demonstrated with the Guardant360 CDx assay in a prospective clinical study involving 282 patients with newly diagnosed metastatic NSCLC71 and in another study involving 1,687 patients with advanced-stage gastrointestinal cancers72, in which more actionable mutations, faster turnaround times and increased patient enrolment in clinical trials were achieved compared with tumour tissue-based testing. Moreover, in a study involving 8,388 patients with advanced-stage NSCLC, alterations were detected in actionable oncogenes in 48% of patients using the Guardant360 assay, with actionable fusions detected in the cfDNA of 2.3% of these patients67. Use of the Guardant360 assay in patients with breast cancer has also revealed that mutations in ESR1 and PTEN are associated with intrinsic resistance to treatment with aromatase inhibitors plus alpelisib73.
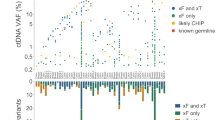
Overall, the feasibility of applying cfDNA assays in the clinical setting, and the ability of such assays to identify common mutations and uncommon gene fusions that can guide the use of molecularly targeted therapies has been well demonstrated74,75. However, whether the use of multigene panels results in improved clinical outcomes across various cancers, as compared with the use of single-gene assays, remains to be determined. Although results obtained with specific multigene cfDNA assays are typically highly concordant with tumour tissue-based assays32,71,76,77, data on the concordance between the results obtained with different multigene plasma cfDNA assays are needed78,79. When replicate plasma samples from 24 patients with different cancers of different stages were used to compare cfDNA variant calls among four commercial NGS gene-panel assays, all assays performed well for somatic alterations with a variant allele freqeuncy (VAF) >10%; however, substantial variability in sensitivity and positive predictive value was observed between assays, which was mostly attributable to technical factors, such as background noise, bioinformatic filtering thresholds and germline variant calls, especially in samples containing cfDNAs with VAFs <1%79.
In August 2020, the Guardant360 CDx assay was approved by the FDA as a comprehensive genomic profiling (CGP) test for the simultaneous assessment of SNVs or indels in 55 tumour-associated genes, CNAs in two genes and fusions in four genes using plasma cfDNA samples. As the first FDA-approved liquid biopsy CGP test, this assay was approved as a companion diagnostic specifically to identify EGFR mutations that predict benefit from osimertinib (exon 19 deletions, L858R in exon 21 or T790M in exon 20) in patients with NSCLC80. As with the aforementioned single-gene assays, a negative cfDNA test result does not necessarily mean that the tumour is negative for these alterations and should, therefore, prompt reflex testing of a tumour tissue biopsy sample using an FDA-approved assay, if feasible. In addition, the Guardant platform was approved as a complementary diagnostic81 for tumour mutation profiling in patients diagnosed with any solid malignancy to provide information that can be used, in conjunction with other laboratory and clinical findings and in accordance with professional guidelines, for clinical decision-making80.
Another CGP liquid biopsy test, F1 Liquid CDx, which interrogates 324 genes, is also approved by the FDA for the detection of substitution mutations and indels in 311 genes, as well as rearrangements in four genes and CNAs in three genes, using plasma cfDNA. This assay was originally approved as a companion diagnostic for use in patients with NSCLC or prostate cancer and has more recently been updated to include additional drugs and use in patients with ovarian or breast cancer. Specifically, this assay is now indicated for use as a companion diagnostic to identify the following groups of patients: patients with NSCLC harbouring EGFR exon 19 deletions or exon 21 L858R mutations who are likely to benefit from gefitinib, erlotinib or osimertinib, as well as ALK rearrangements that predict benefit from alectinib; men with BRCA1/2-mutant or ATM-mutant prostate cancer who are candidates for treatment with olaparib; men with BRCA1/2-mutant prostate cancer and women with BRCA1/2-mutant ovarian cancer who are eligible for treatment with rucaparib; and patients with breast cancer harbouring specific PIK3CA mutations, who could benefit from alpelisib82. Again, patients who might be candidates for these drugs but test negative for the genetic alterations in cfDNA should be reflexed to routine testing of a tumour biopsy sample using an FDA-approved assay, if feasible. Similar to the Guardant360 CDx assay, F1 Liquid CDx is also approved as a complementary diagnostic test to provide tumour mutation profiling of cfDNA in patients with solid tumours, which can be used by qualified health-care professionals in accordance with accepted guidelines. For both of these ctDNA CGP tests, samples are sent to the companies for central processing.
An important question is whether tumour tissue and ctDNA genotyping can be complementary for precision medicine in patients with advanced-stage cancer. Insights into this issue come from a prospective observational study32, in which the FDA-approved MSK-IMPACT NGS panel was used for the analysis of FFPE tumour specimens and the GRAIL 508-gene cfDNA panel was used to evaluate plasma and matched buffy coat samples. Both tissue and cfDNA assay results could be generated for 124 (77%) of 161 patients with advanced-stage cancer (NSCLC, breast cancer or castration-resistant prostate cancer (CRPC)). The cfDNA assay was successfully performed with samples from 151 (94%) of the 161 patients, whereas evaluable data from the tumour tissue analysis were available from 134 patients (84%). The lack of evaluable tumour tissue results in 16% of patients was attributed to insufficient tumour tissue and/or low tumour cellularity, low DNA quantity and/or quality, failed library preparation or other technical issues32. Moreover, some patients in prospective clinical trials lack accessible metastatic lesions, refuse to undergo biopsy sampling or have medical contraindications to biopsy, which can additionally preclude tumour tissue genotyping in around one-third of patients with metastatic cancer, as found in the prospective SAFIR01/UNICANCER trial83.
Thus, cfDNA genotyping is an attractive alternative to tumour tissue profiling, not least because blood can be obtained sequentially and with minimal risk of harm. However, evidence also indicates that plasma ctDNA genotyping using either single-gene PCR assays or multigene NGS panels is not feasible in a proportion of patients with advanced-stage cancer owing to undetectable levels of ctDNA. Even in the aforementioned study of the highly sensitive GRAIL assay32, 20 (16%) of 124 patients with one or more variants detected in tumour tissue had no detectable tumour-derived mutation in their cfDNA. It is well known that ctDNA levels are often low or undetectable in patients with a low tumour burden, cancer at specific sites and specific histologies (such as glioma)84 or tumours that have low levels of proliferation, apoptosis and/or vascularization32,85. Moreover, striking data from the study of the GRAIL assay32 also indicate that >50% of genetic alterations detected in cfDNA from patients with non-hypermutated cancer might be attributable to clonal haematopoiesis of white blood cells, rather than being derived from tumours, which is a phenomenon associated with increasing age86. This phenomenon is termed clonal haematopoiesis of indeterminate potential (CHIP) when detected in individuals with no detectable haematological malignancy87 and is characterized by clonal populations of myeloid cells in the bone marrow or blood that harbour an acquired mutation, often in leukaemia-associated driver genes and other genes (for example, DNMT3A, TET2, ASXL1, JAK2 and TP53, and in more sensitive assays, BRAF, KRAS, NRAS and PIK3CA)88,89,90. Of crucial importance to liquid biopsy is that CHIP can affect the interpretation of cfDNA assay results if unpaired ddPCR or NGS assays are performed, especially when low-VAF ctDNAs are identified90,91. This issue is particularly relevant when determining eligibility for targeted treatments, as further discussed below. In a study of patients with advanced-stage prostate cancer92, ~10% of men had detectable cfDNA harbouring CHIP mutations in DNA repair genes, including ATM, BRCA2 and CHEK2, which could have incorrectly indicated eligibility for poly(ADP ribose) polymerase inhibitors in the absence of paired analyses of DNA from peripheral blood leukocytes. Notably, these CHIP ‘misdiagnoses’ were only detected in men ≥51 years of age.
On the basis of current evidence, one can conclude that plasma ctDNA genotyping can be complementary to tissue genotyping, and vice versa. However, in patients with newly diagnosed advanced-stage cancer, it is not clear whether both tumour tissue and plasma cfDNA analyses using large panels always need to be performed in initial biomarker assessments. In patients with no accessible metastatic lesions to biopsy, plasma cfDNA is an alternative source of material for biomarker identification. In patients with accessible lesions who are eligible for FDA-approved companion diagnostic cfDNA assays, plasma genotyping should be the first option for patient ease and comfort, with reflex tumour tissue analysis reserved only for patients with no detectable targets. In a research setting in which concurrent tissue and plasma genotyping is performed, plasma cfDNA assays can reveal a fraction of mutations that are not identified in tumour specimens76; however, the clinical relevance of these findings to treatment responses remains under investigation.
Clinical validation of ctDNA assays regularly involves and requires large-cohort phase III trials, and these trials increasingly occur within the purview of pharmaceutical companies. Phase III trials now commonly include mandatory blood sampling, although the results from analyses of these samples are often not reported. This paradox has slowed the development of clinically validated blood-based biomarkers and, thus, regulatory approvals of liquid biopsy assays. The development of such biomarkers provides both an important opportunity and, arguably, a responsibility for the developers of such trials. Reporting of liquid biopsy findings should, in the future, be a regular part of the presentation of trial results, and the design of future trials should have statistical analysis plans that include a well-defined biomarker as well as efficacy end points.
Currently, cfDNA assays are only approved as companion diagnostics for a few specific cancers and targeted therapies, as outlined. Outside these indications, screening of many patients would be required to identify a small percentage of patients (around 10%) who could benefit from FDA-approved drugs or be matched to early phase clinical trials of new therapies93. Whether these initial estimates persist will, of course, depend on progress made in translating findings from both prospective clinical trials and real-world evidence databases into new indications for novel or existing molecularly targeted therapies. There is, however, some reason for optimism on this front. As an example, the phase III trial that led to the approval of alpelisib for advanced-stage, PIK3CA-mutant, HR+/HER2− breast cancer demonstrated the utility of ctDNA testing in this setting94, thus launching ctDNA testing into a setting in which ~40% of patients are now eligible for an FDA-approved agent. Similarly, results from the plasmaMATCH trial demonstrate that testing for multiple individually rare pathogenic mutations in patients with advanced-stage breast cancer can collectively generate useful therapeutic ‘hits’ for a substantial percentage of patients (35%)95.
Detection of resistance mechanisms
A major challenge in the application of ctDNA in guiding treatment decisions is the frequent occurrence of co-mutations or CNAs and the development of resistance mutations. Several mechanisms of resistance to targeted therapy can be monitored using plasma ctDNA analysis, including co-mutations that can affect treatment decisions in multiple cancer types, most notably in patients with NSCLC96 and CRC97. For example, KRAS mutation is a mechanism of resistance to EGFR-targeted therapy in patients with CRC that can be detected using plasma cfDNA98,99. cfDNA can also be used to detect, for example, the EGFRT790M resistance mutation in patients with EGFR-mutant NSCLC in order to decide on the optimal subsequent treatment100. Methods to enable detection of CNAs utilizing multigene cfDNA assays, even with low levels of input cfDNA, have been described101. Nevertheless, CNAs are often not detectable using gene panels, and shallow WGS of cfDNA is another tool for detecting such mechanisms of resistance102,103.
In addition to monitoring for known resistance mechanisms, cfDNA can be used to identify unknown mechanisms of treatment resistance. One of the first studies in this area involved the analysis of serial plasma cfDNA samples using WES and provided insights into the mechanisms of resistance to commonly used chemotherapeutic or targeted agents. For example, a patient with metastatic HR+/HER2− breast cancer who progressed following paclitaxel treatment had increased mutant allele fractions of PIK3CA, BMI1 and SMC4 (ref.104). However, WES of plasma cfDNA may only be feasible in patients with a high ctDNA fraction (higher than 5–10%). More targeted approaches have used cfDNA assays including genes with some prior evidence of an association with resistance to a particular drug105. Such a strategy can help validate or refute preclinical hypotheses. As an example, the prevalence of loss of function mutations in RB1 in patients treated with CDK4/6 inhibitors was substantially lower than was expected based on preclinical research105.
However, whether patients with advanced-stage disease need to be evaluated longitudinally using large (>50 gene) cfDNA panels, or whether a targeted approach should be used instead, remains unclear. In contexts in which resistance mechanisms are not known, all patients could be monitored with large gene panels. By contrast, in settings in which resistance mechanisms are well described, follow-up assessments could involve either ddPCR assays for specific mutations or NGS of smaller panels of genes. Thus, ctDNA can be used to probe resistance mechanisms when a valid ‘rescue’ therapy tailored to the resistance mechanism is available (for example, osimertinib for patients with NSCLC and EGFRT790M detected in cfDNA following prior treatment with erlotinib or gefitinib)100.
In terms of efforts to elucidate mechanisms of resistance using cell-based technologies, the CTC-based androgen receptor splice variant 7 (AR-V7) test for metastatic CRPC (mCRPC) represents a success story. AR-V7 is a ligand-independent constitutively active form of the androgen receptor that is not inhibited by anti-androgen therapies, including abiraterone and enzalutamide106. The results of multiple clinical studies indicate that the presence of AR-V7 in CTCs is associated with a poor prognosis in the setting of secondary hormone therapies for mCRPC106,107. Indeed, data from the multicentre prospective PROPHECY study validated the presence of AR-V7+ CTCs, detected using either mRNA or protein assays, as an independent predictor of unfavourable progression-free survival (PFS) and overall survival (OS), after adjusting for CTC number and clinical prognostic factors108.
Other mechanisms of resistance relate to spatial and temporal heterogeneity, including the development of subclones with different resistance mechanisms that are often not all detected in single-tissue biopsy samples, but can be identified using cfDNA109 or CTCs50. In both univariate and multivariate analyses, higher VAFs in cfDNA have been associated with worse outcomes in patients with advanced-stage cancers110. However, as cfDNA measurement technologies and bioinformatics pipelines improve, more low-VAF ctDNAs will be identified; some might reflect CHIP, as discussed previously32, but others might be associated with prognosis or predict drug response. For example, in a study using an ultrasensitive ddPCR assay, the detection of EGFRT790M mutations at VAFs of >1% in tissue sections of EGFR-mutated NSCLCs portended unfavourable PFS and OS with first-generation or second-generation EGFR TKIs, whereas most patients with VAFs of ≤1% had slower disease progression111. Thus, threshold determinations for low-VAF mutations should offer fertile ground for future discoveries.
With regard to CTCs, these analytes are live cells that have, in many patients, resisted previous treatments. Indeed, they are molecularly diverse cells from the primary tumour and/or multiple metastases that might have evolved uniquely over time owing to genetic instability, environmental cues and drug selection pressures, thereby providing a comprehensive snapshot of the tumour biology in real-time112. Thus, single-cell analysis of CTCs can reveal heterogeneity and divergent phenotypes between multiple metastatic tumour sites, providing biological and treatment insights when examined for genomic, transcriptomic, proteomic and functional characteristics50,113,114,115. These insights, in turn, might lead to a better understanding of drug resistance mechanisms and assist in guiding further drug selection112,116.
In addition to targeted therapies, liquid biopsy of ctDNA or CTCs might help in understanding and monitoring resistance to immunotherapy. This topic has been previously reviewed in this journal117 and elsewhere118 and is not developed further here.
Liquid biopsy versus imaging
Pooled analyses of individual patient data from five trials in the setting of mCRPC demonstrated that CTC enumeration using the CELLSEARCH platform can be used as an early measure of treatment response119. Similarly, the clinical validity of CTC enumeration with CELLSEARCH as an adverse prognostic factor has been shown in patients with advanced-stage breast cancer120,121, prostate cancer122 or CRC123; however, CTC enumeration does not inform patient management because no additional clinical utility has been demonstrated compared with standard imaging-based measures. Indeed, although data from the phase III SWOG S0500 trial confirmed the prognostic validity of CTCs, they failed to demonstrate that an early change in chemotherapy based on the persistence of five or more CTCs per 7.5 ml of blood after one cycle of initial chemotherapy can improve OS, as compared with a treatment change upon radiological detection of disease progression124. To many, this result represents not so much a failure of CTC enumeration (which indeed has clear clinical validity in determining PFS and OS) as it does the dangers of tying the development of a technology to inefficacious and unselected treatments. Phenotypic or functional analysis of persistent CTCs to identify actionable aberrations, such as PIK3CA125,126, or new therapeutic targets might be an interesting research avenue to demonstrate clinical utility.
In a proof-of-concept study, tracking patient-specific PIK3CA and TP53 mutations in plasma ctDNA was shown to provide an earlier measure of response to systemic treatment than imaging assessment127. As with CTCs, no data suggest that an early treatment change based on ctDNA-detected as opposed to imaging-detected progression improves clinical outcome in patients with metastatic disease, and thus ctDNA is not currently used for monitoring treatment response.
Liquid biopsy for early stage cancer
Prognostication using CTCs
Historically, data on the prognostic value of CTCs in early stage breast cancer have been obtained through detection of cytokeratin-19 mRNA in the mononuclear cell fraction of peripheral blood128. However, most data come from studies involving a total of a few thousands of patients with breast cancer in whom the CELLSEARCH assay was used to detect CTCs before and/or following surgery and neoadjuvant and/or adjuvant chemotherapy20. These studies demonstrated that patients with detectable CTCs either at diagnosis129,130 or after 5 years of endocrine treatment131 have worse outcomes than those without detectable CTCs. Indeed, CTC detection using CELLSEARCH at baseline could potentially be used as a stratification factor in clinical trials evaluating new drugs.
Nevertheless, several concerns exist regarding the use of CELLSEARCH to detect minimal residual disease (MRD) during follow-up surveillance of patients with early stage breast cancer, not least owing to the low number of CTCs detected in this group132. Another important concern relates to the relatively low sensitivity and specificity of a positive CTC test for predicting disease relapse. In a study involving 1,087 patients with high-risk early stage breast cancer who underwent CTC testing with CELLSEARCH at 2 years after completion of chemotherapy and with a median follow-up duration of 3 years following this test, while the presence of CTCs was prognostic, the sensitivity and specificity of a positive CTC status (defined as one or more CTCs per 7.5 ml of blood) for disease relapse were only 36% and 84%, respectively133. One might argue that serial blood testing and longer follow-up might improve the sensitivity and specificity of the assay, but this hypothesis remains to be proven.
ctDNA-based follow-up assessments
ctDNA levels in patients with early stage disease are usually lower than those in patients with metastatic disease85, which poses substantial challenges for the use of ctDNA surveillance to detect early relapse. One of the first reported studies, albeit in patients with either non-metastatic CRC or CRC metastatic to the liver, focused on the use of ctDNA to predict relapse in 18 patients who had undergone surgery (either colectomy or hepatic metastasectomy) with curative intent134. After tumour DNA sequencing, patient-specific mutations were evaluated in plasma cfDNA at 13−56 days following surgery. The half-life of ctDNA after surgery was estimated to be 114 min. Moreover, 13 of 14 patients with the detection of ctDNA (93%) had disease relapse, generally within a year, in contrast to none of four patients without detectable ctDNA134, indicating its potential role in the measurement of tumour dynamics. However, only two of those patients had non-metastatic CRC, and long-term follow-up data are lacking; thus, conclusions could not be drawn about the use of ctDNA as a biomarker in the setting of early stage disease. Following a proof-of-concept study in 55 patients135, Garcia-Murillas et al.136 found evidence of the prognostic value of ctDNA analysis during follow-up surveillance in a cohort of 170 patients with early stage breast cancer who had received neoadjuvant and/or adjuvant chemotherapy. The investigators used ddPCR assays to monitor patient-specific mutations (one mutation only in the majority of patients) in plasma cfDNA that had been previously identified through targeted NGS of primary tumour tissue. Personalized ctDNA assays were successfully developed for 101 (60%) of 170 patients; in the remaining patients, no mutation could be identified in the primary tumour. In this study, detection of ctDNA was associated with an increased risk of disease relapse, with a hazard ratio of 25.2 (95% CI 6.7–95.6; P < 0.001) at a median follow-up duration of 35.5 months. Interestingly, TP53 mutations in three patients were identified as being due to CHIP; these patients remained relapse-free136.
Of note, different cancers have a different somatic mutation frequency137, which can affect the use of ctDNA assays for follow-up surveillance of early stage cancers after standard local and systemic treatment. Another challenge in the implementation of ctDNA for surveillance of early stage cancer is the low fraction of ctDNA in cfDNA and consequently the presence of tumour mutations in plasma at VAFs potentially below the background sequencing error threshold. The development of methods to overcome such limitations might improve analytical sensitivity138,139,140. Another approach for addressing limited abundance of cfDNA is the use of WGS to increase the breadth of sequencing141.
The blood volume analysed is another important consideration when aiming to detect low-abundance ctDNAs. An elegant modelling demonstrated that to be able to detect de novo a single mutation with a VAF of 0.01% with 95% confidence would require 150–300 ml with 30,000× sequencing coverage142. However, the blood sample volume required to detect any of ten known mutations with VAFs of 0.01% is the same as that required to detect a single 0.1% VAF mutation tested in isolation. Thus, the sensitivity of a given ctDNA assay in patients with localized cancers is dependent on not only the blood volume analysed, but also the number of mutations screened143. Commercially available platforms incorporating this strategy of probing multiple known mutations via personalized ctDNA assays have been developed85,144. For example, the Signatera assay, which was designated a Breakthough Device by the FDA in 2019, involves the selection of 16 somatic variants identified through WES of paired primary tumour and germline DNA samples, followed by the design of patient-specific assays using multiplex-PCR amplification and subsequently NGS85. In a retrospective case–control study involving 49 patients with breast cancer, including 18 with disease relapse, a mean of about four plasma samples per patient were analysed, and the sensitivity and specificity of the Signatera assay in the prediction of disease relapse were 89% and 100%, respectively144. Another approach for personalized tracking of mutations, called targeted digital sequencing (TARDIS), has been developed using patient-specific primer panels and clinically relevant blood volumes (one to two tubes of blood) to detect ctDNA at the very low concentrations expected in patients undergoing systemic treatment for non-metastatic cancer. When used to monitor response to neoadjuvant chemotherapy, ctDNA concentrations were 5.7-fold lower in patients with early stage breast cancer who had a pathological complete response (pCR) (median allele fraction 0.003%) than in those who had residual disease (median allele fraction 0.018%)143.
The prognostic role of ctDNA detected using various assays has been evaluated in over 800 patients with early stage breast cancer across eight retrospective studies and one prospective trial (Table 2). Data from four retrospective studies demonstrate a median lead time from ctDNA detection to radiological relapse of up to 11 months in patients who do not receive additional systemic treatment. The promising results of ctDNA-based prognostication mostly come from single-centre studies; therefore, data from additional prospective, multicentre series with longer follow-up durations, and/or meta-analyses of reported studies, are needed to confirm the sensitivity, specificity and lead time to relapse.
Treating ctDNA relapse
Here, we introduce the term ‘ctDNA relapse’ to describe a disease stage in which patients present with detectable ctDNA during routine cancer surveillance but without overt imaging-detected disease relapse after completion of surgery and neoadjuvant and/or adjuvant chemotherapy for their primary cancer. Systemic treatment for ctDNA relapse could potentially create a new setting for drug testing beyond the metastatic, adjuvant, neoadjuvant and post-neoadjuvant settings that have been traditionally used in clinical trials (Table 3).
Historically, cancer drugs (such as trastuzumab, a monoclonal antibody targeting the HER2 receptor on breast tumour cells) have first been tested and approved in the metastatic setting145 followed by studies in the neoadjuvant and/or adjuvant settings146,147,148. More recently, promising novel agents have been evaluated in patients who have high risk of disease relapse based on the absence of a pCR after standard neoadjuvant-based treatment (known as the post-neoadjuvant setting). For example, the anti-HER2 antibody–drug conjugate trastuzumab emtansine (T-DM1) was approved by the FDA for the treatment of patients with HER2+ breast cancer and no pCR after HER2-targeted neoadjuvant treatment149. Absence of pCR is useful to identify those patients who have a high risk of disease relapse, particularly those with triple-negative or HER2-positive breast cancer, whereas the prognostic value of pCR in patients with HR+/HER2− breast cancer is less clear150. One of the reasons for this disparity is that the subsequent adjuvant endocrine treatment has an impact on the association between pCR and long-term outcomes. Moreover, patients with HR+/HER2− breast cancer can have disease relapse 20 years or more after diagnosis151; thus, real-time biomarkers are needed to identify those at high risk of relapse. ctDNA detection during follow-up surveillance could potentially provide such a real-time biomarker. Notably, evaluation of different systemic treatments, such as CDK4/6 inhibitors, in this ctDNA relapse setting is planned152.
Currently, a crucial question relating to this new setting remains: can systemic treatment of ctDNA relapse lead to cure, or will such treatment simply delay the development of overt (radiologically or clinically detectable) metastatic disease without improving cure rates (Box 1)? Various possible clinical trial designs can be envisioned for treatments aiming to improve the outcomes of patients with such ctDNA relapse of non-metastatic cancer (Fig. 3). For example, patients could be randomly assigned either to an experimental arm involving ctDNA profiling and subsequent allocation of biomarker-defined experimental therapies or to a standard-of-care (SoC) control arm using only primary tumour histology-based treatments without ctDNA profiling (Fig. 3, Design 1). Alternatively, patients could first be stratified according to ctDNA status (positive or negative), with each group subsequently randomized to experimental or SoC treatment groups (Design 2). Other designs focus on ctDNA-positive patients only and involve either a randomized comparison of experimental versus SoC treatment (Design 3) or a single-arm evaluation of a new experimental treatment (Design 4). Design 1 and Design 2 would require the highest number of patients but provide the highest level of evidence for the clinical utility of ctDNA as a biomarker. With Designs 1, 2 and 3, the primary objective would usually be to compare invasive disease-free survival (iDFS) between the experimental and SoC treatment arms. For Design 2, this comparison will be performed both in the ctDNA-positive and ctDNA-negative patients, whereas for Design 3 it will be performed only in ctDNA-positive patients. Design 1 would mostly likely be used in a non-inferiority trial, whereas Designs 2 and 3 would typically be applied in a superiority trial. Designs 3 and 4 require fewer patients and are more appropriate for ctDNA assays with robust evidence of clinical validity and in settings in which ctDNA detection has been demonstrated to have a high sensitivity and specificity for predicting disease relapse. With Designs 1, 2, 3 and 4, associations between early ctDNA elimination (for example, after one treatment cycle) and iDFS can be evaluated in patients who are ctDNA-positive at study entry. Designs 1, 2, 3 and 4 can lead to treatment escalation for ctDNA-positive patients. Designs 2, 3, and 4 are focused on late adjuvant therapy and patients who present with ctDNA relapse during follow-up monitoring (for example, patients with oestrogen receptor-positive breast cancer), but could also apply to the post-neoadjuvant setting in patients with a substantial residual cancer burden at the time of surgery and evidence of ctDNA or CTCs in blood following surgical resection.
Design 1 enables the comparison of primary tumour-guided versus circulating tumour DNA (ctDNA)-guided adjuvant treatment. This approach involves ctDNA analysis for the detection of minimal residual disease (MRD) following initial surgery (or other curative-intent treatments) and is predicated on early application of adjuvant therapy for those with detectable MRD who have a high risk of disease relapse (for example, in patients with stage II colorectal cancer, in whom ctDNA detection after curative surgery is being investigated as a means to decide whether to administer adjuvant chemotherapy). Design 2 involves the comparison of an experimental therapy versus a standard-of-care treatment in both ctDNA-positive and ctDNA-negative groups. Design 3 compares an experimental versus a standard-of-care treatment only in the ctDNA-positive group. Design 4 evaluates an experimental adjuvant treatment in the ctDNA-positive group only (single-arm study). Design 5 explores the value of treating patients at ctDNA relapse according to different druggable molecular aberrations detected in either plasma or tissue. These trial designs can also be applied with any other liquid biopsy-based biomarker, including circulating tumour cells (CTCs). cfDNA, cell-free DNA.
Finally, an alternative design (Fig. 3, Design 5) involves patients at ctDNA relapse who would be treated according to the druggable molecular aberrations detected in their cfDNA. This design might be feasible in the future when the technology becomes available for reliable and CGP of plasma cfDNA, including the ability to detect low abundance ctDNAs. Alternatively, this approach could be used today through CGP of a patient’s primary cancer tissue followed by monitoring for the druggable molecular aberration in plasma.
Beyond designing trials focused on treating patients at ctDNA relapse, the value of ctDNA detection shortly after definitive surgery, as a marker of MRD, in guiding decisions on the use of adjuvant therapy is currently being tested. For example, this strategy is being evaluated in patients with stage II colon cancer in the Circulate trial153 and the E-Dynamic trial154.
Before the current efforts to treat ctDNA relapse152, investigators tried in the past to target MRD155 (for example, detected as disseminated tumour cells in bone marrow156 or CTCs in peripheral blood157) using delayed adjuvant treatment strategies; however, to date, none of these efforts has resulted in an approved clinical indication. Successful examples of treating patients with high-risk disease before imaging-detected relapse include the hormone therapies apalutamide, enzalutamide and darolutamide, which have all been approved by the FDA for the treatment of patients with non-metastatic CRPC and rising serum prostate-specific antigen (PSA) levels on the basis of improvements in metastasis-free survival observed in placebo-controlled phase III trials158,159,160. With regard to haematological malignancies, an example of the successful treatment of MRD is provided by the approval of blinatumomab for the treatment of patients with B cell precursor acute lymphoblastic leukaemia and MRD after initial multiagent chemotherapy. This approval was based on data from an open-label, single-arm study (consistent with Design 4, Fig. 3) involving 116 patients, which showed that those with a complete MRD response after one cycle of blinatumomab had longer PFS and OS durations than non-responders161.
Liquid biopsy for early cancer detection
In addition to challenges related to low ctDNA levels that are associated with early stage cancers, the low incidence of cancer in the general population is an important challenge to the use of liquid biopsy — as well as other screening methodologies — for cancer diagnosis. Nevertheless, the potential to screen for cancer using blood samples is attractive, and several liquid biopsy approaches for cancer diagnosis are under development. Initial approaches have been based on the detection of driver gene mutations in plasma cfDNA162. However, this approach might be hampered by the presence of CHIP-related mutations in a substantial proportion of individuals without cancer and, moreover, might require the detection of genomic aberrations specific to certain tumour types, thus limiting the diagnostic scope of the assay13,163.
Another approach is based on combined analyses of circulating proteins and cancer-associated mutations in plasma, such as the CancerSeek platform164. Other approaches are predicated on the analysis of epigenetic alterations that might be tissue-specific and cancer type-specific by analysing genome-wide differentially methylated regions via cell-free methylated DNA immunoprecipitation and high-throughput bisulfite-free sequencing (cfMeDIP-seq)165 or other methylation patterns166. Alternative approaches include the analysis of differences in genome-wide fragmentation patterns between non-tumour and tumour cfDNA using the DNA evaluation of fragments for early interception (DELFI) platform167 or by fragment size168; cfDNA fragmentation patterns have also been used to infer differential accessibility of transcription factor binding sites that are associated with certain cancers169. More recently, targeted methylation analysis of samples from the Circulating Cell-free Genome Atlas (CCGA) study and the STRIVE study has enabled the development of a methylation-based assay for the simultaneous detection and tissue-of-origin identification of various cancers across disease stages170.
An example of the potential clinical utility of a liquid biopsy approach for early detection has been provided by the DETECT-A study171. In this prospective study, 10,006 women aged 65–75 years with no prior history of cancer were evaluated using the CancerSeek platform. Women who tested positive by a liquid biopsy baseline test with exclusion of CHIP (n = 490, 4.9%) then underwent a second confirmatory test (n = 134, 1.3%) followed by a diagnostic PET–CT scan. Overall, 26 women had cancers that were detected using CancerSeek, including five with stage I (19%), three with stage II (12%), eight with stage III (31%) and nine with stage IV (35%) cancers, as well as one with cancer of unknown stage but without metastases. Of these 26 women, 14 (54%) had ctDNA mutations, 11 (42%) had increased levels of protein biomarkers and one (4%) had both. During the study period, however, 70 of a total of 96 (73%) cancers were detected by standard-of-care screening (n = 24, 25%) or because of symptoms or by other means (n = 46, 48%). These results, while interesting, arguably indicate that this particular approach is likely to have limited clinical utility, given that most cancers were detected by other means and that 17 of the 26 (65%) cancers detected by the blood test were diagnosed at an advanced stage. Thus, further refinement of this test is ongoing. Whether this approach would improve clinical outcomes in an unselected population remains unknown in the absence of a randomized controlled trial.
A detailed description of the different liquid biopsy assays for cancer diagnosis and the ongoing clinical studies designed to validate these assays is beyond the scope of this Perspective and have been addressed elsewhere74,172. However, we highlight several considerations that need to be taken into account during the clinical development of a blood-based or any other diagnostic test for cancer.
False-positives and false-negatives
Even a small percentage of false-positive test results, spread across a national population, would hugely increase the demand for confirmatory imaging as well as biopsy sampling of imaging-detected benign abnormalities. Thus, false-positives have obvious implications for health-care resources as well as patient well-being. Conversely, false-negative results would have important implications related to delays in diagnosis.
Effects on overall survival
The assumption that early cancer diagnosis leads to improved OS is not necessarily correct171. Certainly, early diagnosis of some cancer types might not improve survival outcomes; for example, when metastatic disease is readily curable with systemic therapy (as is the case for many testicular cancers) or, conversely, when the cancer will still prove to be incurable owing to a lack of effective systemic therapy (for example, limited-stage small-cell lung cancer). Additionally, an early stage cancer that is detected via a liquid biopsy might be indolent or would never have developed into a life-threatening cancer (for example, many prostate and breast cancers), thus constituting ‘overdiagnosis’ of incidental cancers that are unlikely to affect a patient’s overall health or lifespan. More specifically, early detection of prostate cancer through serum PSA testing of men aged 55–69 years carries a grade of ‘C’ from the US Preventive Services Task Force. This recommendation means that the screening test should be offered selectively based on professional judgment and patient preference because the net benefit in reducing the risk of death from prostate cancer is small and the potential harms due to false-positive results, overdiagnosis and overtreatment (such as complications that include incontinence or erectile dysfunction) are considerable173.
Psychological effects
The availability of a liquid biopsy approach for population-level early cancer detection would potentially alter or redefine what it means to be a ‘patient’ or to ‘have a disease’. Some otherwise healthy, asymptomatic people will be delighted to have that disease identified at an early stage, but for many such a diagnosis will constitute an unwelcome and potentially devastating event. Arguably, diagnostic testing is only valuable if it leads to action that can alter the disease outcome, particularly by defining an appropriate (and preferably life-saving) therapeutic strategy.
Clinical integration of liquid biopsy
Considering the various current or potential future clinical applications of liquid biopsy, and assuming that the assay is demonstrated to have appropriate analytical and clinical validity, clinical utility, and approval and reimbursement for a particular indication, the actual integration of liquid biopsy tests into the clinical workflow is an important and complex aspect that is often overlooked. To better understand this process, experience can be drawn from successful examples of tissue-based multigene assays that are currently used in the management of patients with cancer, such as Oncotype Dx and FoundationOne CDx (F1CDx). Oncotype Dx was developed to identify a subset of patients with HR+, node-negative breast cancer who can be spared chemotherapy174, whereas F1CDx was developed to identify druggable genomic aberrations in patients with advanced-stage cancers175. These assays are both performed at a central laboratory and have changed the paradigm of cancer diagnostics from one that was historically based on tumour analysis in the pathology department of each hospital (decentralized testing) to a model that is based on off-site, centralized testing (Table 4). This paradigm shift reflects the fact that multigene assays, as opposed to single-gene assays, require considerable investment of time, knowledge and resources, not only in their development but also in their day-to-day clinical implementation. This investment encompasses state-of-the-art NGS machines, adequate cloud storage space for the sequencing data, technicians for wet laboratory tasks and bioinformaticians for data analysis, as well as a molecular board of clinicians, biologists and bioinformaticians for data interpretation. Central laboratories performing these assays are providing their services at a national or even international level, which potentially enables cost savings through economies of scale (that is, owing to the large number of samples analysed). Many local pathology laboratories will be unable to compete with central laboratories when it comes to the implementation of multigene assays. We predict that the model of decentralized testing will be used for single-gene or small multigene (fewer than five genes) ctDNA assays, but centralized testing will prevail for larger multigene ctDNA assays.
Another important aspect for the incorporation of liquid biopsy into the clinical workflow involves the education and training of hospital personnel, including pathologists, nurses and physicians, in the application and interpretation of the new assay. For example, even at a local hospital where multigene tumour and/or liquid biopsy assays will not be performed on-site, but rather in a central laboratory, a nurse or certified phlebotomist is required to perform the blood draw and the pathology laboratory personnel perform the initial sample processing, if needed; thus, individuals in these roles have to be aware of the assay protocols and any preanalytical issues that might affect assay performance. Similarly, the physician who will receive the assay report needs training in how to read the report, interpret the results and act accordingly. Such training for hospital personnel needs to be organized through dedicated seminars delivered at work or, even earlier, to be included in the university curriculum as part of education in new technologies. Despite the crucial role of training of individual laboratory and clinical physicians in the fundamental principles of liquid biopsy assays, the rapid accumulation of knowledge, occurring at warp speed across many tumour types, makes it difficult for most physicians to keep up to date. In many health-care systems, this situation has led to the development of multidisciplinary molecular tumour boards charged with the interpretation of results obtained by NGS of ctDNA or tumour-tissue DNA176. Such tumour boards typically incorporate clinical, molecular genetic and pathology expertise, and increasingly offer support to medical oncologists grappling with often confusing and complex information overload. Going forwards, harmonization of the approaches used by molecular tumour boards will provide important opportunities to increase efficiency and clinical use of liquid biopsy assays.
AI to facilitate liquid biopsy
Artificial intelligence (AI) promises to revolutionize the way we practise medicine177. AI has already been leveraged to improve the performance of different liquid biopsy assays and will facilitate their future integration into the clinical workflow. Examples include the use of machine learning approaches for the detection and characterization of CTCs178,179,180, for the analysis of ctDNA for cancer detection and localization165,167,181, for integrative multi-omics analyses182 and future integration of liquid biopsy tests together with other clinicogenomic, metabolomic, immunomic, microbiomic and homeostatic data to guide treatment decisions.
With regard to early diagnosis, a machine learning platform termed ‘lung cancer likelihood in plasma’ (Lung-CLiP) has been developed for lung cancer detection based on targeted sequencing of plasma cfDNA and matched leukocyte DNA183. The Lung-CLiP algorithm was trained using samples from patients with well-annotated clinical, histological and imaging data, particularly metabolic tumour volume (MTV) data from PET–CT scans. In retrospective analyses, Lung-CLiP distinguished patients with early stage NSCLC from control individuals with matched risk profiles, with a sensitivity of 41%, 54% and 67% for stage I, II and III disease, respectively, when the specificity was set at 98%, and was also validated in an independent prospectively obtained cohort. Moreover, it was shown that the sensitivity of Lung-CLiP strongly correlated with MTV, with approximate sensitivities of 16% for a volume of 1 ml, 52% for 10 ml, and 80% for >100 ml (ref.183).
Priorities for liquid biopsy research
Outlining the key current challenges in liquid biopsy might reveal priorities for future research in this area. Here, we provide our perspective on the top ten challenges (Box 2).
Standardization of the preanalytical variables previously discussed is of the utmost importance, given that such variables can cause the results of the same assay to differ substantially and can ultimately lead to problems in clinical interpretation. Standardization of the crucial procedures before the actual performance of the liquid biopsy assay, together with adequate reporting of these procedures, will not only improve the value of the assays in guiding treatment decisions but also enable the comparison and/or combination of results from different studies. As highlighted, the use of AI is expected to improve the performance of liquid biopsy assays and accelerate their introduction into clinical practice, and therefore demands increased attention. Relatedly, data sharing is essential. International academic–industry collaborations in genomics and in oncology184 are aiming to accelerate research in these fields, and we advocate for such ventures in liquid biopsy research. As an often-overlooked area, the incorporation of liquid biopsy assays into the clinical workflow warrants additional studies.
In terms of clinical decision-making, another key challenge relates to the role of aberrations detected in plasma cfDNA but not in tumour tissue and their relevance to treatment selection. For a given somatic aberration that is known to be predictive of benefit from a molecularly targeted agent, whether patients with this aberration in plasma cfDNA but not in synchronous tumour tissue derive the same benefit as patients who have the aberration detected in both plasma cfDNA and tumour tissue or in tumour tissue only is now beginning to be studied185. This question could be addressed either in retrospective series or optimally in prospective clinical trials.
Another challenge lies in demonstrating the value of CTCs as a complementary tool to ctDNA in precision medicine. ctDNA can be used for real-time monitoring of the evolution of the tumour genome, while CTCs can provide complementary information about changes to the transcriptome and proteome. The combined use of distinct and diverse data from ctDNA and CTC analyses might, therefore, refine our ability to predict benefit from a particular targeted agent. With continued technology development, and perhaps in concert with organ-on-a-chip platforms that could potentially recapitulate appropriate microenvironmental contexts, we postulate that CTCs might also be used for real-time drug screening. Clinical trials are needed to evaluate these possible roles of CTCs, especially with regard to specific targeted treatments.
Whether scope exists for liquid biopsy biomarkers (CTCs or ctDNA) to complement standard imaging assessment by CT in monitoring treatment response is another pertinent question. We believe that CTCs or ctDNA might have such a role186. For example, we can envisage utility in monitoring metastatic disease that is evaluable but non-measurable by standard imaging, such as bone-only metastatic breast cancer. CTCs or ctDNA responses could potentially also be used instead of or in addition to CT assessments, at the same time points, to provide more accurate assessments of treatment response, for example, in confirming pseudoprogression with immune-checkpoint inhibitors117. Notably, CTC and/or ctDNA assays might be less costly than CT or PET–CT imaging, are unlikely to cause secondary cancers, are less bothersome to patients and are more likely to provide insights into mechanisms of treatment resistance. Well-conducted studies with analytically and clinically validated liquid biopsy assays are needed to demonstrate clinical utility in disease monitoring instead of or in combination with standard imaging. Such studies will require the development of rigorous and validated definitions of CTC or ctDNA responses, akin to Response Evaluation Criteria in Solid Tumors (RECIST) used for radiographical response assessments187.
Efforts to improve the OS of patients with early stage cancers through the timely detection and treatment of ctDNA relapse present additional challenges. Currently, whether treating ctDNA relapse would lead to cure or simply delay the development of imaging-detected metastatic disease is unknown (Box 1); the answer to this question might differ according to the disease setting and the treatments used.
Finally, the use of liquid biopsy for early cancer detection will be challenging — but potentially invaluable. Randomized controlled trials comparing the use of ctDNA assays (either with or without prior screening tools) with standard diagnostic algorithms are needed to assess effects on OS, as well as the quality of life of those undergoing these liquid biopsy tests. Similar trials that evaluated previous generation technologies (such as screening mammography188 and serum PSA testing189) provide a template for the clinical testing of liquid biopsy-based screening.
Conclusions
In this Perspective, we outline several challenges in liquid biopsy research, including scientific questions related not only to technology development and clinical research but also to the optimal integration of liquid biopsy into the clinical workflow, an aspect that is often neglected in the literature. Addressing these challenges in the coming years will lead to a profound change in the practice of oncology by introducing liquid biopsy as a tool for the real-time assessment and management of tumour evolution.
References
McGranahan, N. & Swanton, C. Clonal heterogeneity and tumor evolution: past, present, and the future. Cell 168, 613–628 (2017).
López, S. et al. Interplay between whole-genome doubling and the accumulation of deleterious alterations in cancer evolution. Nat. Genet. 52, 283–293 (2020).
Yates, L. R. & Campbell, P. J. Evolution of the cancer genome. Nat. Rev. Genet. 13, 795–806 (2012).
Russo, M. et al. Adaptive mutability of colorectal cancers in response to targeted therapies. Science 366, 1473–1480 (2019).
Heitzer, E., Haque, I. S., Roberts, C. E. S. & Speicher, M. R. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat. Rev. Genet. 20, 71–88 (2019).
Hudson, T. J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010).
Hoadley, K. A. et al. Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173, 291–304.e6 (2018).
ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 578, 82–93 (2020).
Rozenblatt-Rosen, O. et al. The Human Tumor Atlas Network: charting tumor transitions across space and time at single-cell resolution. Cell 181, 236–249 (2020).
Navin, N. et al. Tumour evolution inferred by single-cell sequencing. Nature 472, 90–94 (2011).
Ramalingam, N. & Jeffrey, S. S. Future of liquid biopsies with growing technological and bioinformatics studies: opportunities and challenges in discovering tumor heterogeneity with single-cell level analysis. Cancer J. 24, 104–108 (2018).
Bertucci, F. et al. Genomic characterization of metastatic breast cancers. Nature 569, 560–564 (2019).
Reiter, J. G. et al. An analysis of genetic heterogeneity in untreated cancers. Nat. Rev. Cancer 19, 639–650 (2019).
Hanahan, D. & Coussens, L. M. Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21, 309–322 (2012).
Pantel, K. & Alix-Panabières, C. Circulating tumour cells in cancer patients: challenges and perspectives. Trends Mol. Med. 16, 398–406 (2010).
Wan, J. C. M. et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer 17, 223–238 (2017).
Cherry, S. R. et al. Total-body PET: maximizing sensitivity to create new opportunities for clinical research and patient care. J. Nucl. Med. 59, 3–12 (2018).
Ignatiadis, M., Lee, M. & Jeffrey, S. S. Circulating tumor cells and circulating tumor DNA: challenges and opportunities on the path to clinical utility. Clin. Cancer Res. 21, 4786–4800 (2015).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Pantel, K. & Alix-Panabières, C. Liquid biopsy and minimal residual disease — latest advances and implications for cure. Nat. Rev. Clin. Oncol. 16, 409–424 (2019).
Cescon, D. W., Bratman, S. V., Chan, S. M. & Siu, L. L. Circulating tumor DNA and liquid biopsy in oncology. Nat. Cancer 1, 276–290 (2020).
Lee, J. S., Park, S. S., Lee, Y. K., Norton, J. A. & Jeffrey, S. S. Liquid biopsy in pancreatic ductal adenocarcinoma: current status of circulating tumor cells and circulating tumor DNA. Mol. Oncol. 13, 1623–1650 (2019).
Teutsch, S. M. et al. The Evaluation of Genomic Applications in Practice and Prevention (EGAPP) initiative: methods of the EGAPP working group. Genet. Med. 11, 3–14 (2009).
Cheng, D. T. et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J. Mol. Diagn. 17, 251–264 (2015).
Zehir, A. et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat. Med. 23, 703–713 (2017).
Samstein, R. M. et al. Tumor mutational load predicts survival after immunotherapy across multiple cancer types. Nat. Genet. 51, 202–206 (2019).
US Food and Drug Administration. MSK-IMPACT (Integrated Mutation Profiling of Actionable Cancer Targets). FDA https://www.accessdata.fda.gov/cdrh_docs/pdf17/DEN170058.pdf (2017).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 2017, PO.17.00011 (2017).
New York State Department of Health, Wadsworth Center. Memorial Hosp For Cancer and Allied Diseases Dept of Pathology. New York State https://www.wadsworth.org/memorial-hosp-for-cancer-and-allied-diseases-dept-of-pathology-115 (2020).
Razavi, P. et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat. Med. 25, 1928–1937 (2019).
Mateo, J. et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO Scale for Clinical Actionability of Molecular Targets (ESCAT). Ann. Oncol. 29, 1895–1902 (2018).
Katsoulakis, E., Duffy, J. E., Hintze, B., Spector, N. L. & Kelley, M. J. Comparison of annotation services for next-generation sequencing in a large-scale precision oncology program. JCO Precis. Oncol. 4, 212–221 (2020).
Wagner, A. H. et al. A harmonized meta-knowledgebase of clinical interpretations of somatic genomic variants in cancer. Nat. Genet. 52, 448–457 (2020).
Van Norman, G. A. Drugs and devices: comparison of European and US approval processes. JACC Basic. Transl. Sci. 1, 399–412 (2016).
Pittella-Silva, F. et al. Plasma or serum: which is preferable for mutation detection in liquid biopsy? Clin. Chem. 66, 946–957 (2020).
Parpart-Li, S. et al. The effect of preservative and temperature on the analysis of circulating tumor DNA. Clin. Cancer Res. 23, 2471–2477 (2017).
Gerber, T. et al. Assessment of pre-analytical sample handling conditions for comprehensive liquid biopsy analysis. J. Mol. Diagn. 22, 1070–1086 (2020).
Salvianti, F. et al. The pre-analytical phase of the liquid biopsy. New Biotechnol. 55, 19–29 (2020).
Grölz, D. et al. Liquid biopsy preservation solutions for standardized pre-analytical workflows — venous whole blood and plasma. Curr. Pathobiol. Rep. 6, 275–286 (2018).
Van Paemel, R. et al. Genome-wide study of the effect of blood collection tubes on the cell-free DNA methylome. Epigenetics https://doi.org/10.1080/15592294.2020.1827714 (2020).
Greytak, S. R. et al. Harmonizing cell-free DNA collection and processing practices through evidence-based guidance. Clin. Cancer Res. 26, 3104–3109 (2020).
Merker, J. D. et al. Circulating tumor DNA analysis in patients with cancer: American Society of Clinical Oncology and College of American Pathologists joint review. J. Clin. Oncol. 36, 1631–1641 (2018).
Andree, K. C., van Dalum, G. & Terstappen, L. W. M. M. Challenges in circulating tumor cell detection by the CellSearch system. Mol. Oncol. 10, 395–407 (2016).
Wong, K. H. K. et al. Whole blood stabilization for the microfluidic isolation and molecular characterization of circulating tumor cells. Nat. Commun. 8, 1733 (2017).
Hodgkinson, C. L. et al. Tumorigenicity and genetic profiling of circulating tumor cells in small-cell lung cancer. Nat. Med. 20, 897–903 (2014).
Yu, M. et al. Ex vivo culture of circulating breast tumor cells for individualized testing of drug susceptibility. Science 345, 216–220 (2014).
Fehm, T. N. et al. Diagnostic leukapheresis for CTC analysis in breast cancer patients: CTC frequency, clinical experiences and recommendations for standardized reporting. Cytometry A 93, 1213–1219 (2018).
Keller, L. & Pantel, K. Unravelling tumour heterogeneity by single-cell profiling of circulating tumour cells. Nat. Rev. Cancer 19, 553–567 (2019).
Cook, L. et al. Does size matter? Comparison of extraction yields for different-sized DNA fragments by seven different routine and four new circulating cell-free extraction methods. J. Clin. Microbiol. 56, e01061-18 (2018).
Meddeb, R., Pisareva, E. & Thierry, A. Guidelines for the preanalytical conditions for analyzing circulating cell-free DNA. Clin. Chem. 65, 623–633 (2019).
Lampignano, R. et al. Multicenter evaluation of circulating cell-free DNA extraction and downstream analyses for the development of standardized (pre)analytical work flows. Clin. Chem. 66, 149–160 (2020).
Febbo, P. G. et al. Minimum technical data elements for liquid biopsy data submitted to public databases. Clin. Pharmacol. Ther. 107, 730–734 (2020).
Foundation for the National Institutes of Health. Biomarkers Consortium – Identification and validation of ctDNA quality control materials. FNIH https://fnih.org/ctdna (2020).
Hao, Y. X. et al. Effectiveness of circulating tumor DNA for detection of KRAS gene mutations in colorectal cancer patients: a meta-analysis. Onco. Targets Ther. 10, 945–953 (2017).
Oxnard, G. R. et al. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin. Cancer Res. 20, 1698–1705 (2014).
Bidard, F.-C. et al. Circulating tumor cells and circulating tumor DNA detection in potentially resectable metastatic colorectal cancer: a prospective ancillary study to the Unicancer PRODIGE-14 trial. Cells 8, 516 (2019).
US Food and Drug Administration. cobas® EGFR Mutation Test v2. FDA https://www.accessdata.fda.gov/cdrh_docs/pdf15/P150047A.pdf (2016).
André, F. et al. Alpelisib for PIK3CA-mutated, hormone receptor-positive advanced breast cancer. N. Engl. J. Med. 380, 1929–1940 (2019).
US Food and Drug Administration. therascreen PIK3CA RGQ PCR kit. FDA https://www.accessdata.fda.gov/cdrh_docs/pdf19/P190004A.pdf (2019).
Kumar, S. et al. Tracking plasma DNA mutation dynamics in estrogen receptor positive metastatic breast cancer with dPCR-SEQ. NPJ Breast Cancer 4, 39 (2018).
Sabari, J. K. et al. A prospective study of circulating tumor DNA to guide matched targeted therapy in lung cancers. J. Natl Cancer Inst. 111, 575–583 (2019).
Clark, T. A. et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J. Mol. Diagn. 20, 686–702 (2018).
Zill, O. A. et al. The landscape of actionable genomic alterations in cell-free circulating tumor DNA from 21,807 advanced cancer patients. Clin. Cancer Res. 24, 3528–3538 (2018).
Aggarwal, C. et al. Clinical implications of plasma-based genotyping with the delivery of personalized therapy in metastatic non-small cell lung cancer. JAMA Oncol. 5, 173–180 (2019).
Mack, P. C. et al. Spectrum of driver mutations and clinical impact of circulating tumor DNA analysis in non-small cell lung cancer: analysis of over 8000 cases. Cancer 126, 3219–3228 (2020).
Pritchett, M. A. et al. Prospective clinical validation of the InVisionFirst-Lung circulating tumor DNA assay for molecular profiling of patients with advanced nonsquamous non-small-cell lung cancer. JCO Precis. Oncol. 3, PO.18.00299 (2019).
Finzel, A., Sadik, H., Ghitti, G. & Laes, J.-F. The combined analysis of solid and liquid biopsies provides additional clinical information to improve patient care. J. Cancer Metastasis Treat. 4, 21 (2018).
Owonikoko, T. K. et al. Randomized phase II study of paclitaxel plus alisertib versus paclitaxel plus placebo as second-line therapy for SCLC: primary and correlative biomarker analyses. J. Thorac. Oncol. 15, 274–287 (2020).
Leighl, N. B. et al. Clinical utility of comprehensive cell-free DNA analysis to identify genomic biomarkers in patients with newly diagnosed metastatic non-small cell lung cancer. Clin. Cancer Res. 25, 4691–4700 (2019).
Nakamura, Y. et al. Clinical utility of circulating tumor DNA sequencing in advanced gastrointestinal cancer: SCRUM-Japan GI-SCREEN and GOZILA studies. Nat. Med. 26, 1859–1864 (2020).
Razavi, P. et al. Alterations in PTEN and ESR1 promote clinical resistance to alpelisib plus aromatase inhibitors. Nat. Cancer 1, 382–393 (2020).
Kilgour, E., Rothwell, D. G., Brady, G. & Dive, C. Liquid biopsy-based biomarkers of treatment response and resistance. Cancer Cell 37, 485–495 (2020).
Drilon, A. et al. Antitumor activity of crizotinib in lung cancers harboring a MET exon 14 alteration. Nat. Med. 26, 47–51 (2020).
Rothé, F. et al. Plasma circulating tumor DNA as an alternative to metastatic biopsies for mutational analysis in breast cancer. Ann. Oncol. 25, 1959–1965 (2014).
Bachet, J. B. et al. RAS mutation analysis in circulating tumor DNA from patients with metastatic colorectal cancer: the AGEO RASANC prospective multicenter study. Ann. Oncol. 29, 1211–1219 (2018).
Torga, G. & Pienta, K. J. Patient-paired sample congruence between 2 commercial liquid biopsy tests. JAMA Oncol. 4, 868–870 (2018).
Stetson, D. et al. Orthogonal comparison of four plasma NGS tests with tumor suggests technical factors are a major source of assay discordance. JCO Precis. Oncol. 3, 1–9 (2019).
US Food and Drug Administration. Guardant360® CDx. FDA https://www.accessdata.fda.gov/cdrh_docs/pdf20/P200010A.pdf (2020).
Scheerens, H. et al. Current status of companion and complementary diagnostics: strategic considerations for development and launch. Clin. Transl. Sci. 10, 84–92 (2017).
US Food and Drug Administration. FoundationOne® Liquid CDx (F1 Liquid CDx). FDA https://www.accessdata.fda.gov/cdrh_docs/pdf20/P200016A.pdf (2020).
André, F. et al. Comparative genomic hybridisation array and DNA sequencing to direct treatment of metastatic breast cancer: a multicentre, prospective trial (SAFIR01/UNICANCER). Lancet Oncol. 15, 267–274 (2014).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 6, 224ra24 (2014).
Abbosh, C. et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 545, 446–451 (2017).
Shlush, L. I. Age-related clonal hematopoiesis. Blood 131, 496–504 (2018).
Steensma, D. P. et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 126, 9–16 (2015).
Steensma, D. P. & Ebert, B. L. Clonal hematopoiesis as a model for premalignant changes during aging. Exp. Hematol. 83, 48–56 (2020).
Acuna-Hidalgo, R. et al. Ultra-sensitive sequencing identifies high prevalence of clonal hematopoiesis-associated mutations throughout adult life. Am. J. Hum. Genet. 101, 50–64 (2017).
Hu, Y. et al. False-positive plasma genotyping due to clonal hematopoiesis. Clin. Cancer Res. 24, 4437–4443 (2018).
Chan, H. T., Chin, Y. M., Nakamura, Y. & Low, S. K. Clonal hematopoiesis in liquid biopsy: from biological noise to valuable clinical implications. Cancers 12, 2277 (2020).
Jensen, K. et al. Association of clonal hematopoiesis in DNA repair genes with prostate cancer plasma cell-free DNA testing interference. JAMA Oncol. https://doi.org/10.1001/jamaoncol.2020.5161 (2020).
Rothwell, D. G. et al. Utility of ctDNA to support patient selection for early phase clinical trials: the TARGET study. Nat. Med. 25, 738–743 (2019).
Juric, D. et al. Alpelisib + fulvestrant for advanced breast cancer: subgroup analyses from the phase III SOLAR-1 trial [abstract]. Cancer Res 79 (Suppl. 4), GS3-08 (2019).
Turner, N. C. et al. Circulating tumour DNA analysis to direct therapy in advanced breast cancer (plasmaMATCH): a multicentre, multicohort, phase 2a, platform trial. Lancet Oncol. 21, 1296–1308 (2020).
Li, S. et al. Coexistence of EGFR with KRAS, or BRAF, or PIK3CA somatic mutations in lung cancer: a comprehensive mutation profiling from 5125 Chinese cohorts. Br. J. Cancer. 110, 2812–2820 (2014).
De Roock, W., De Vriendt, V., Normanno, N., Ciardiello, F. & Tejpar, S. KRAS, BRAF, PIK3CA, and PTEN mutations: implications for targeted therapies in metastatic colorectal cancer. Lancet Oncol. 12, 594–603 (2011).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536 (2012).
Diaz, L. A. et al. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature 486, 537–540 (2012).
Mok, T. S. et al. Osimertinib or platinum-pemetrexed in EGFR T790M-positive lung cancer. N. Engl. J. Med. 376, 629–640 (2017).
Li, B. T. et al. Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: results from the actionable genome consortium. Ann. Oncol. 30, 597–603 (2019).
Heitzer, E. et al. Tumor-associated copy number changes in the circulation of patients with prostate cancer identified through whole-genome sequencing. Genome Med. 5, 30 (2013).
Bourrier, C. et al. Shallow whole-genome sequencing from plasma identifies FGFR1 amplified breast cancers and predicts overall survival. Cancers 12, 1481 (2020).
Murtaza, M. et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature 497, 108–112 (2013).
O’Leary, B. et al. The genetic landscape and clonal evolution of breast cancer resistance to palbociclib plus fulvestrant in the PALOMA-3 trial. Cancer Discov. 8, 1390–1403 (2018).
Antonarakis, E. S. et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N. Engl. J. Med. 371, 1028–1038 (2014).
Scher, H. I. et al. Assessment of the validity of nuclear-localized androgen receptor splice variant 7 in circulating tumor cells as a predictive biomarker for castration-resistant prostate cancer. JAMA Oncol. 4, 1179–1186 (2018).
Armstrong, A. J. et al. Prospective multicenter validation of androgen receptor splice variant 7 and hormone therapy resistance in high-risk castration-resistant prostate cancer: the PROPHECY study. in. J. Clin. Oncol. 37, 1120–1129 (2019).
Parikh, A. R. et al. Liquid versus tissue biopsy for detecting acquired resistance and tumor heterogeneity in gastrointestinal cancers. Nat. Med. 25, 1415–1421 (2019).
Pairawan, S. et al. Cell-free circulating tumor DNA variant allele frequency associates with survival in metastatic cancer. Clin. Cancer Res. 26, 1924–1931 (2020).
Beau-Faller, M. et al. Independent prognostic value of ultra-sensitive quantification of tumor pre-treatment T790M subclones in EGFR mutated non-small cell lung cancer (NSCLC) treated by first/second generation TKI, depends on variant allele frequency (VAF): results of the French Cooperative Thoracic Intergroup (IFCT) Biomarkers France project. Lung Cancer 140, 19–26 (2020).
Ma, N. & Jeffrey, S. S. Deciphering cancer clues from blood. Science 367, 1424–1425 (2020).
Ignatiadis, M. et al. Prognostic value of the molecular detection of circulating tumor cells using a multimarker reverse transcription-PCR assay for cytokeratin 19, mammaglobin A, and HER2 in early breast cancer. Clin. Cancer Res. 14, 2593–2600 (2008).
Powell, A. A. et al. Single cell profiling of circulating tumor cells: transcriptional heterogeneity and diversity from breast cancer cell lines. PLoS ONE 7, e33788 (2012).
Lim, S. et al. Liquid biopsy: one cell at a time. NPJ Precis. Oncol. 3, 23 (2019).
Ebright, R. Y. et al. Deregulation of ribosomal protein expression and translation promotes breast cancer metastasis. Science 367, 1468–1473 (2020).
Cabel, L. et al. Clinical potential of circulating tumour DNA in patients receiving anticancer immunotherapy. Nat. Rev. Clin. Oncol. 15, 639–650 (2018).
Hofman, P., Heeke, S., Alix-Panabières, C. & Pantel, K. Liquid biopsy in the era of immuno-oncology: is it ready for prime-time use for cancer patients? Ann. Oncol. 30, 1448–1459 (2019).
Heller, G. et al. Circulating tumor cell number as a response measure of prolonged survival for metastatic castration-resistant prostate cancer: a comparison with prostate-specific antigen across five randomized phase III clinical trials. J. Clin. Oncol. 36, 572–580 (2018).
Cristofanilli, M. et al. Circulating tumor cells, disease progression, and survival in metastatic breast cancer. N. Engl. J. Med. 351, 781–791 (2004).
Bidard, F.-C. et al. Clinical validity of circulating tumour cells in patients with metastatic breast cancer: a pooled analysis of individual patient data. Lancet Oncol. 15, 406–414 (2014).
Moreno, J. G. et al. Circulating tumor cells predict survival in patients with metastatic prostate cancer. Urology 65, 713–718 (2005).
Cohen, S. J. et al. Prognostic significance of circulating tumor cells in patients with metastatic colorectal cancer. Ann. Oncol. 20, 1223–1229 (2009).
Smerage, J. B. et al. Circulating tumor cells and response to chemotherapy in metastatic breast cancer: SWOG S0500. J. Clin. Oncol. 32, 3483–3489 (2014).
Markou, A. et al. PIK3CA mutational status in circulating tumor cells can change during disease recurrence or progression in patients with breast cancer. Clin. Cancer Res. 20, 5823–5834 (2014).
Gasch, C. et al. Frequent detection of PIK3CA mutations in single circulating tumor cells of patients suffering from HER2-negative metastatic breast cancer. Mol. Oncol. 10, 1330–1343 (2016).
Dawson, S. J. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 368, 1199–1209 (2013).
Ignatiadis, M. et al. Different prognostic value of cytokeratin-19 mRNA-positive circulating tumor cells according to estrogen receptor and HER2 status in early-stage breast cancer. J. Clin. Oncol. 25, 5194–5202 (2007).
Riethdorf, S. et al. Prognostic impact of circulating tumor cells for breast cancer patients treated in the neoadjuvant “Geparquattro” trial. Clin. Cancer Res. 23, 5384–5393 (2017).
Bidard, F.-C. et al. Circulating tumor cells in breast cancer patients treated by neoadjuvant chemotherapy: a meta-analysis. J. Natl Cancer Inst. 110, 560–567 (2018).
Sparano, J. et al. Association of circulating tumor cells with late recurrence of estrogen receptor-positive breast cancer: a secondary analysis of a randomized clinical trial. JAMA Oncol. 4, 1700–1706 (2018).
Thery, L. et al. Circulating tumor cells in early breast cancer. JNCI Cancer Spectr. 3, pkz026 (2019).
Trapp, E. et al. Presence of circulating tumor cells in high-risk early breast cancer during follow-up and prognosis. J. Natl Cancer Inst. 111, 380–387 (2019).
Diehl, F. et al. Circulating mutant DNA to assess tumor dynamics. Nat. Med. 14, 985–990 (2008).
Garcia-Murillas, I. et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci. Transl. Med. 7, 302ra133 (2015).
Garcia-Murillas, I. et al. Assessment of molecular relapse detection in early-stage breast cancer. JAMA Oncol. 5, 1473–1478 (2019).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and search for new cancer genes. Nature 499, 214–218 (2013).
Berger, M. F. & Mardis, E. R. The emerging clinical relevance of genomics in cancer medicine. Nat. Rev. Clin. Oncol. 15, 353–365 (2018).
Newman, A. M. et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat. Biotechnol. 34, 547–555 (2016).
Wang, T. T. et al. High efficiency error suppression for accurate detection of low-frequency variants. Nucleic Acids Res. 47, e87 (2019).
Zviran, A. et al. Genome-wide cell-free DNA mutational integration enables ultra-sensitive cancer monitoring. Nat. Med. 26, 1114–1124 (2020).
Haque, I. S. & Elemento, O. Challenges in using ctDNA to achieve early detection of cancer. Preprint at bioRxiv https://doi.org/10.1101/237578 (2017).
McDonald, B. R. et al. Personalized circulating tumor DNA analysis to detect residual disease after neoadjuvant therapy in breast cancer. Sci. Transl. Med. 11, eaax7392 (2019).
Coombes, R. C. et al. Personalized detection of circulating tumor DNA antedates breast cancer metastatic recurrence. Clin. Cancer Res. 25, 4255–4263 (2019).
Slamon, D. J. et al. Use of chemotherapy plus a monoclonal antibody against HER2 for metastatic breast cancer that overexpresses HER2. N. Engl. J. Med. 344, 783–792 (2001).
Piccart-Gebhart, M. J. et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N. Engl. J. Med. 353, 1659–1672 (2005).
Romond, E. H. et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N. Engl. J. Med. 353, 1673–1684 (2005).
Gianni, L. et al. Neoadjuvant chemotherapy with trastuzumab followed by adjuvant trastuzumab versus neoadjuvant chemotherapy alone, in patients with HER2-positive locally advanced breast cancer (the NOAH trial): a randomised controlled superiority trial with a parallel HER-negative cohort. Lancet 375, 377–384 (2010).
Von Minckwitz, G. et al. Trastuzumab emtansine for residual invasive HER2-positive breast cancer. N. Engl. J. Med. 380, 617–628 (2019).
Cortazar, P. et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet 384, 164–172 (2014).
Pan, H. et al. 20-year risks of breast-cancer recurrence after stopping endocrine therapy at 5 years. N. Engl. J. Med. 377, 1836–1846 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT04567420 (2020).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT04089631 (2020).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03826758 (2020).
Pantel, K. & Hayes, D. F. Disseminated breast tumour cells: biological and clinical meaning. Nat. Rev. Clin. Oncol. 15, 129–131 (2018).
Naume, B. et al. Clinical outcome with correlation to disseminated tumor cell (DTC) status after DTC-guided secondary adjuvant treatment with docetaxel in early breast cancer. J. Clin. Oncol. 32, 3848–3857 (2014).
Ignatiadis, M. et al. Trastuzumab versus observation for HER2 nonamplified early breast cancer with circulating tumor cells (EORTC 90091-10093, BIG 1-12, Treat CTC): a randomized phase II trial. Ann. Oncol. 29, 1777–1783 (2018).
Smith, M. R. et al. Apalutamide treatment and metastasis-free survival in prostate cancer. N. Engl. J. Med. 378, 1408–1418 (2018).
Hussain, M. et al. Enzalutamide in men with nonmetastatic, castration-resistant prostate cancer. N. Engl. J. Med. 378, 2465–2474 (2018).
Fizazi, K. et al. Darolutamide in nonmetastatic, castration-resistant prostate cancer. N. Engl. J. Med. 380, 1235–1246 (2019).
Gökbuget, N. et al. Blinatumomab for minimal residual disease in adults with B-cell precursor acute lymphoblastic leukemia. Blood 131, 1522–1531 (2018).
Phallen, J. et al. Direct detection of early-stage cancers using circulating tumor DNA. Sci. Transl. Med. 9, eaan2415 (2017).
Bick, A. G. et al. Inherited causes of clonal haematopoiesis in 97,691 whole genomes. Nature 586, 763–768 (2020).
Cohen, J. D. et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 359, 926–930 (2018).
Shen, S. Y. et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature 563, 579–583 (2018).
Hao, X. et al. DNA methylation markers for diagnosis and prognosis of common cancers. Proc. Natl Acad. Sci. USA 114, 7414–7419 (2017).
Cristiano, S. et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature 570, 385–389 (2019).
Mouliere, F. et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci. Transl. Med. 10, eaat4921 (2018).
Ulz, P. et al. Inference of transcription factor binding from cell-free DNA enables tumor subtype prediction and early detection. Nat. Commun. 10, 4666 (2019).
Liu, M. C. et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann. Oncol. 31, 745–759 (2020).
Lennon, A. M. et al. Feasibility of blood testing combined with PET-CT to screen for cancer and guide intervention. Science 369, eabb9601 (2020).
Keller, L., Belloum, Y., Wikman, H. & Pantel, K. Clinical relevance of blood-based ctDNA analysis: mutation detection and beyond. Br. J. Cancer https://doi.org/10.1038/s41416-020-01047-5 (2020).
US Preventive Services Task Force. Prostate cancer screening. USPSTF https://www.uspreventiveservicestaskforce.org/uspstf/recommendation/prostate-cancer-screening (2018).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 351, 2817–2826 (2004).
Frampton, G. M. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat. Biotechnol. 31, 1023–1031 (2013).
Van De Haar, J., Hoes, L. & Voest, E. Advancing molecular tumour boards: highly needed to maximise the impact of precision medicine. ESMO Open 4, e000516 (2019).
Topol, E. J. High-performance medicine: the convergence of human and artificial intelligence. Nat. Med. 25, 44–56 (2019).
Zeune, L. L. et al. How to agree on a CTC: evaluating the consensus in circulating tumor cell scoring. Cytometry A 93, 1202–1206 (2018).
Iyer, A. et al. Integrative analysis and machine learning based characterization of single circulating tumor cells. J. Clin. Med. 9, 1206 (2020).
Ota, S. et al. Ghost cytometry. Science 360, 1246–1251 (2018).
Wan, N. et al. Machine learning enables detection of early-stage colorectal cancer by whole-genome sequencing of plasma cell-free DNA. BMC Cancer 19, 832 (2019).
Camacho, D. M., Collins, K. M., Powers, R. K., Costello, J. C. & Collins, J. J. Next-generation machine learning for biological networks. Cell 173, 1581–1592 (2018).
Chabon, J. J. et al. Integrating genomic features for non-invasive early lung cancer detection. Nature 580, 245–251 (2020).
Menetski, J. P. et al. The Foundation for the National Institutes of Health Biomarkers Consortium: past accomplishments and new strategic direction. Clin. Pharmacol. Ther. 105, 829–843 (2019).
Xu, H. et al. A comparison of EGFR mutation status in tissue and plasma cell-free DNA detected by ADx-ARMS in advanced lung adenocarcinoma patients. Transl. Lung Cancer Res. 8, 135–143 (2019).
Budd, G. T. et al. Circulating tumor cells versus imaging–predicting overall survival in metastatic breast cancer. Clin. Cancer Res. 12, 6403–6409 (2006).
Eisenhauer, E. A. et al. New Response Evaluation Criteria in Solid Tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Nelson, H. D. et al. Effectiveness of breast cancer screening: systematic review and meta-analysis to update the 2009 US Preventive Services Task Force recommendation. Ann. Intern. Med. 164, 244–255 (2016).
Ilic, D. et al. Prostate cancer screening with prostate-specific antigen (PSA) test: a systematic review and meta-analysis. BMJ 362, k3519 (2018).
Olsson, E. et al. Serial monitoring of circulating tumor DNA in patients with primary breast cancer for detection of occult metastatic disease. EMBO Mol. Med. 7, 1034–1047 (2015).
Riva, F. et al. Patient-specific circulating tumor DNA detection during neoadjuvant chemotherapy in triple-negative breast cancer. Clin. Chem. 63, 691–699 (2017).
Chen, Y. H. et al. Next-generation sequencing of circulating tumor DNA to predict recurrence in triple-negative breast cancer patients with residual disease after neoadjuvant chemotherapy. NPJ Breast Cancer 3, 24 (2017).
Rothé, F. et al. Circulating tumor DNA in HER2-amplified breast cancer: a translational research substudy of the NeoALTTO phase III trial. Clin. Cancer Res. 25, 3581–3588 (2019).
Zhang, X. et al. Parallel analyses of somatic mutations in plasma circulating tumor DNA (ctDNA) and matched tumor tissues in early-stage breast cancer. Clin. Cancer Res. 25, 6546–6553 (2019).
Radovich, M. et al. Association of circulating tumor DNA and circulating tumor cells after neoadjuvant chemotherapy with disease recurrence in patients with triple-negative breast cancer: preplanned secondary analysis of the BRE12-158 randomized clinical trial. JAMA Oncol. 6, 1410–1415 (2020).
Scher, H. I. et al. Association of AR-V7 on circulating tumor cells as a treatment-specific biomarker with outcomes and survival in castration-resistant prostate cancer. JAMA Oncol. 2, 1441–1449 (2016).
Acknowledgements
The work of M.I. is supported by Les Amis de Bordet and Fondation Contre le Cancer. The work of G.W.S. is supported in part by the Susan G. Komen Foundation. The work of S.S.J. is supported in part by the John and Marva Warnock Research Fund, Natalie and Vladimir Ermakoff, and the Stanford Catalyst for Collaborative Solutions, Stanford School of Engineering.
Author information
Authors and Affiliations
Contributions
All authors contributed to all aspects of manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
M.I. has received consultancy fees from Celgene, Novartis, Pfizer, Seattle Genetics and Tesaro, and travel grants from Amgen and Pfizer. The institution of M.I. has received research grants from Menarini Silicon Biosystems and Natera. G.W.S. is a member of the Board of Directors of Tessa Therapeutics and of the Scientific Advisory Boards of Syndax and Verseau Therapeutics. S.S.J. serves as a scientific advisor for Quantumcyte and Ravel Biotechnology.
Additional information
Peer review information
Nature Reviews Clinical Oncology thanks R. Rosell; F.-C. Bidard, who co-reviewed with L. Cabel; and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Blood Profiling Atlas in Cancer (BloodPAC): https://www.bloodpac.org/
CANCER-ID: https://www.cancer-id.eu/
cBioPortal: https://www.cbioportal.org/index.do
European Liquid Biopsy Society (ELBS): http://www.elbs.eu
OncoKB: https://www.oncokb.org/
OncoPrint: https://bioc.ism.ac.jp/packages/3.2/bioc/vignettes/ComplexHeatmap/inst/doc/s8.oncoprint.html#toc_0
Rights and permissions
About this article
Cite this article
Ignatiadis, M., Sledge, G.W. & Jeffrey, S.S. Liquid biopsy enters the clinic — implementation issues and future challenges. Nat Rev Clin Oncol 18, 297–312 (2021). https://doi.org/10.1038/s41571-020-00457-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41571-020-00457-x
This article is cited by
-
Optical nanomaterial-based detection of biomarkers in liquid biopsy
Journal of Hematology & Oncology (2024)
-
Soluble Periostin is a potential surveillance biomarker for early and long-term response to chemotherapy in advanced breast cancer
Cancer Cell International (2024)
-
Integrating cfDNA liquid biopsy and organoid-based drug screening reveals PI3K signaling as a promising therapeutic target in colorectal cancer
Journal of Translational Medicine (2024)
-
Decoding the glycoproteome: a new frontier for biomarker discovery in cancer
Journal of Hematology & Oncology (2024)
-
Liquid biopsy in T-cell lymphoma: biomarker detection techniques and clinical application
Molecular Cancer (2024)